Structure-based design of key amino acid substitutions to improve enzyme function.

Utilizing machine learning to accelerate the directed evolutions of enzymes with novel functions.

Enzyme biocatalysis offers sustainable and efficient pathways for drug discovery and production.
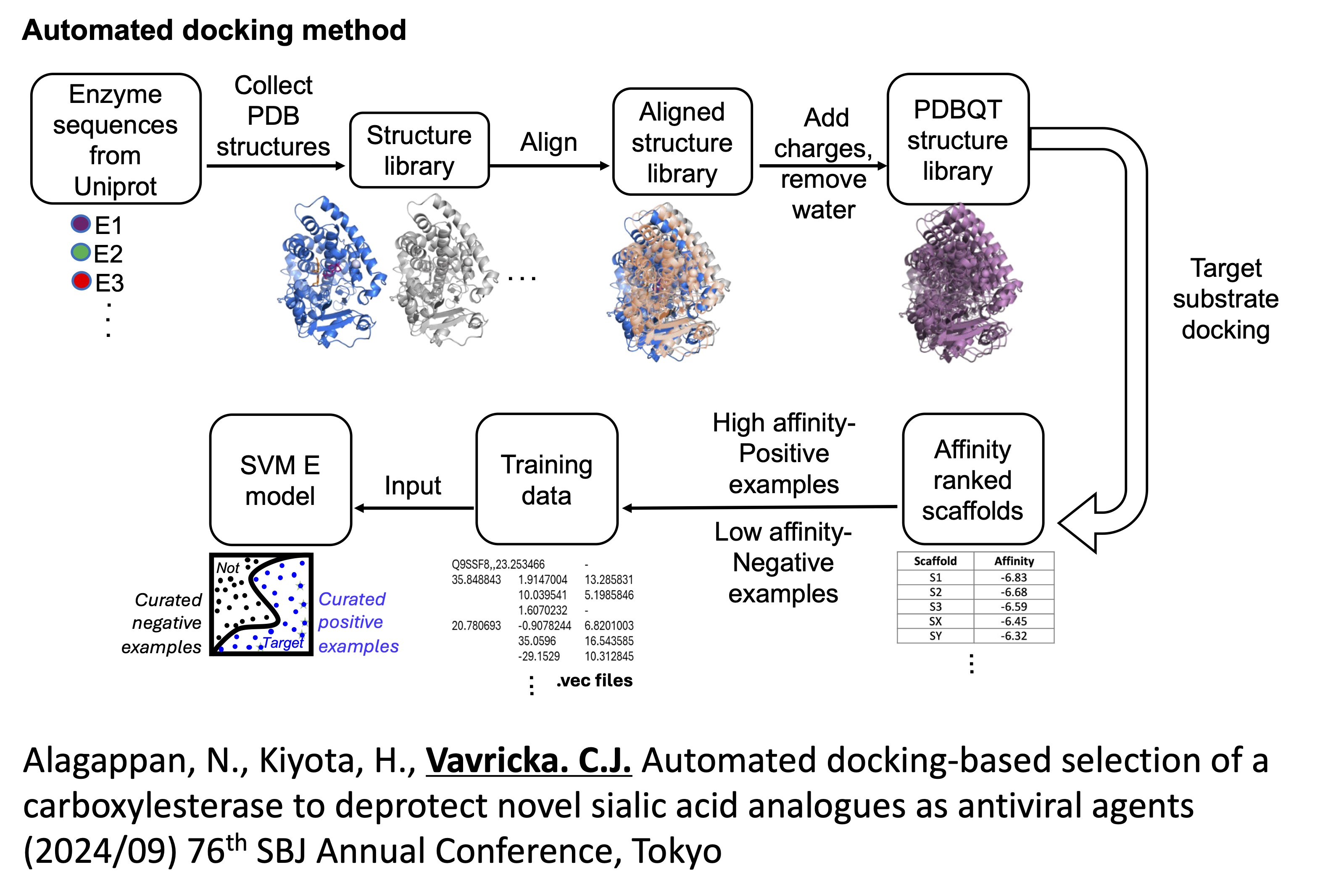
Our research group has recently developed a novel phosphonate analogue of sialic acid as an anti-viral neuraminidase inhibitor. However, deprotection of our key phosphonate ester intermediate using synthetic organic chemistry methods has been proved to be difficult. Therefore, carboxylesterase is explored as a scaffold with potential to cleave phosphonate esters, as well as other carboxylate ester prodrugs. We are selecting optimal scaffolds using an automated docking protocol that screens the binding of our target substrate to the active sites of several hundred carboxylesterases. Then, to further optimize scaffold selection, the automated docking method is utilized to curate training sequences for machine learning-based classification and regression analyses. Based on this workflow, optimal carboxylesterases with high potential to deprotect our target substrate can be automatically selected.